Moving from ST.25 to ST.26 (The New Sequence Listing Standard) – Webinar
Table of Contents
Key-points Covered in the Webinar (Moving from ST. 25 to ST. 26 – The New Sequence Listing Standard)
- What is Sequence Listing Standard and why is it required?
- What are ST.25 and ST.26 standards?
- How does ST.26 differ from ST.25?
- What are the tools and processes involved in preparing ST.26 Sequence Listing?
- How can professionals help in preparing error free Sequence Listing?
Anchor
Devika Saini, Manager LSC, Sagacious IP
Key-note Speaker
Pooja Chhikara, Assistant Manager LSC, Sagacious IP
"*" indicates required fields

Webinar Transcript
Devica is speaking – Welcome everyone to the webinar. This is Davikasani, manager, Life Sciences and Chemistry team at Sagacious, signing in from India to welcome you all to the Webinar. Today, the topic of the Webinar is moving from St 25 to St 26, the new sequence listing standard.
Before I go on to introduce this topic, I am delighted to welcome all the participants from different regions, including the US. Europe, India, China and other Asian countries. Your participation is a wonderful encouragement to the efforts and attempt that we are making to raise awareness and spread knowledge that has been honed by Sagacious over several years of working with inventors are in the organization’s IP departments and IP job practices.
Speaker’s Introduction
So here along with me, I have my colleague, Pooja Chhikara, assistant manager, life sciences and Chemistry team. She’s a qualified patent agent, having more than six years of experience with competency in identifying and assessing IT in biotechnology, microbio, biochemistry, genetic engineering, pharma and immunology and she has managed more than 60 sequence listing projects monthly.
Welcome to the webinar Pooja.
Pooja is speaking – Hey, Davika, thanks for having me on the Webinar.
Devica is speaking – Pleasure.
Initial Remarks – Pooja Chhikara
Pooja Chhikara is speaking – Well, Devica, a lot of patent filings happen in the biotechnology domain wherein biological sequences form an important part of biotech inventions. During patent filings, these biological sequences need to be submitted in specific formats. And recently, WIPO has changed these standards for filing the sequence listings.
Hence, I believe it is pertinent to know and discuss the differences between the new and old sequence listing standard. So that is what we are going to focus today.
Agenda of the Webinar
Devica is speaking – Before we move ahead, I invite our listeners to keep sharing their questions as and when they have it during the course of the session. They can also share their questions via chat box on Presentation window.
We will pick up on those questions and answer them after finishing our brief talk.So, apart from this, the participants can also drop us an email later at Webinar@sagaciousresearch.com and we will answer to that.So now let’s get started with the main part of our presentation. And for that, let me invite Pooja to start with the Webinar. Over to you, Pooja.
Pooja is speaking : Thank you, Devica. So last year we conducted our first webinar on Sequence listing that covered the following topics:
- What is Sequence Listing?
- Patent Office requirements
- Advantages of sequence listing,
- Tools for preparing them
- The key considerations during filing or drafting
- Methodology for preparing normal sequence listings and modified sequences.
So, for more details you can access the webinar recording from our Sagacious website. I have provided it in the PPT also, which you can refer to when the PPT is shared later. So, today’s webinar is a continuation of the last Webinar. However, we will cover few of the previous topics in brief that are necessary for understanding today’s topic.
What is Sequence Listing Standard?
As we know, biotechnology inventions involve gene mutation, gene sequencing and regulating, expressing or silencing of genes. Further, a gene is a chain of polynucleotides which are composed of nucleotide bases like adenine, thiamine, Quenin or encytocene that code for amino acids. So during filing such patent applications that comprise the biological sequences, all the DNA and protein sequences need to be entered into the patent applications and are required to have a particular structure instead of a raw format. So we can say that sequence listing is a list of biological sequences provided in the format prescribed by patent offices.
Moving forward, what does St 25 and St 26 stand for? These are the sequence listing standard or guidelines prescribed by WIPO. The WIPO sequence listing standard, St 25, is fully equivalent to NXC of the Pct administrative instructions that defines requirements for submission of sequence listings. Then, the new sequence listing is also developed by a committee on WIPO sequence listing standard.
Shortcomings of St 25
- First is it is not compliant with INSTC requirements, that is, international nucleotide sequence database collaboration, so the data is lost when entered into public databases.
- Secondly, St 25 rules are not clear and IP offices worldwide interpret and enforce the rules differently.
- Third, the sequence types that are common today that are not covered by St 25 rules, for example, the nucleotide analogs d amino acids branched sequences and therefore are not present in searchable databases.
- Lastly, data is unstructured in St 25, which means St 25 format is difficult to use for automated validation and data exchange.
What’s new about Sequence listing Standard?
Currently, we follow St 25 sequence listing standard to prepare our sequence listings. However, the 54th session of the WIPO General Assembly has approved the new Big bank implementation date of WIPO standard St 26th July 1, 2022 At national, regional and international levels, all intellectual property offices will transition simultaneously at the international, national and regional levels coming to St 26.

Benefits & Advantages of St 26
The advantages of St 26 include:
- Enhancing the accuracy and quality of presentation of sequences for easier dissemination, benefiting applicants, public and examiners.
- It also facilitates searching of the sequence data and allows sequence data to be exchanged in electronic form and to be introduced into computerized databases.
- Next, the standard serves as guidance to ensure agreement among all IP offices on application of sequence rules.
- Further, it allows acceptance of a single sequence listing worldwide except for required translations of language.
- SD 26 increases automation of data validation and streamlined processing by IP offices and it enhances submission quality due to the structure of XML sequence listings.
Sequence Listing Standard – Differences between ST 25 Standard and ST 26 Standard
Let’s discuss these differences in detail. The first difference is based on the file format. The St 25 accept ASI text format and St 26 accept XML file format. Then less than ten specifically defined nucleotides and less than four specifically defined amino acids are permitted in case of St 25, but the same is not permitted under SD 26.
In case of St 25, we can name all applicant and inventors. However, only one applicant and optionally one inventor may be included in case of St 26.
The other differences based on annotation, the St 25 provides us with feature keys only, whereas St 26 provides us with feature keys and qualifiers.
We will see these differences in detail in our next slide. In case of DM IO assets, branched sequences and analogs, they are not required to be included under St 25, but they must be included under St 26. Next difference is based on priority application. All priority application information can be included in case
of St 25, but only the earliest priority application can be included in case of St 26.
Difference based on Title
The next difference is based on title. One invention title is permitted in St 25, but multiple invention titles are permitted in St 26, each one in a different language. In case of applicant or inventor names, they must be in basic Latin characters under St 25, but under St 26 they may be included using any valid Unicode character along with Latin translation.
Further, the St 25 specifies the sequences as DNA, RNA or PRT only, whereas in case of St 26 they keep DNA and RNAs same, but they have replaced the Prtsae or amino acid along with a mandatory mold type qualifier to further describe the molecule. For example, in case of RNA we can further describe them as genomic, RNA, mRNA, tRNA and so on.
On the basis of Uracil symbol
Then the next difference is based on uracil symbol. You represent uracil in nucleotide sequences in case of St 25. However, T represents uracil in RNA sequences and thymine in DNA sequences. Now, symbol is not used in any of the cases.
Difference on the basis of Organism Name
Next difference is based on organism name. If we see here the artificial sequence term of St 25 is replaced with synthetic construct. The unknown term is replaced with unidentified. Next difference is based on variables. The N and Xaa variables must have a definition provided in a feature in case of St 25, but St26 provides default values for the two variables. Presentation of amino acid sequences is done by three letter abbreviations in case of St 25, but the representation is done by one letter abbreviation under St 26.
Then we have mixed mode sequences. Mixed mode sequences are permitted under St 35, wherein nucleotide sequence with amino acid translations are shown below. That is a form of representation, whereas no mixed mode nucleotide translations are included under St 26.
Our next differentiating feature is featured location. In case of St 25, the feature location format was not clearly defined, but now they have defined it very clearly under St 26 by subdividing them under few terms like join order, complement in case of nucleotide sequence.
Features applicable to all sequences
This slide talks about features that are applicable to all sequences, nucleotide as well as amino acid sequences.
- The first feature is about numbering of sequences. Each sequence should be assigned a separate sequence identification number.The identification number must begin with number one and increase consecutively by integers where no sequence is present. For a sequence identification number that is, an intentionally skipped sequence represented as triple zero must be used in place of a sequence.
- Now, next feature is about presentation of sequences.Sequence listing can be divided into two parts a general information part and a sequence data part. And the sequence listing must be presented as a single file in XML using document type definition. So the general information part comprises bibliographic information solely for association of the sequence listing of the patent application for which the sequence listing is submitted.
- Next, the sequence data part is composed of one or more sequence data elements, each of which contain information about one sequence.
further, the sequence data element elements must include feature keys and subsequent qualifiers based on the INSTC that is international Nucleotide Sequence Database Collaboration and Uniprot specifications.

Nucleotide Sequences
Nucleotide
Now we will talk about the presentation of nucleotide sequences. Specifically, a nucleotide sequence must be represented only by a single stand in the five prime to three prime direction from left to right or in the direction from left to right that mimics the five prime to three prime direction. The designation five prime to three prime or any other similar designations must not be included in the sequence.
Now, coming to the numbering part, for the purpose of the standard, the first nucleotide presented in the sequences residue position number one. And when the nucleotide sequences are circular in configuration, applicant must choose the nucleotide and residue position number one and the last residue position number must equal the number of nucleotides in the sequence.
Now, while presenting the ambiguity symbol, the most appropriate symbol, the most restrictive symbol should be used. Like for example, if a nucleotide in a given position could be A or G, then R should be used rather than N.
The symbol N will be construed as any one of ACG or T. This is about presentation of modified and nucleotide sequences.
Modified Nucleotides
Modified nucleotides should be represented in this sequence as the corresponding unmodified nucleotides wherever possible. And when any modified nucleotide in a sequence cannot be represented by any other symbol, then they must be represented by the symbol N. And this symbol N is the equivalent of only one residue. Like for example, here we have mentioned N in place of a modified nucleotide, which could not be represented by any other symbols.
Further, a modified nucleotide must be described in the feature table using the feature key.
We’ll discuss this in detail next slide. So the format used to represent nucleotides include all lower cases symbols and any symbol used to represent the nucleotide is the equivalent of only one residue. No spaces, no numbering is required, no U symbol to be used, t represents uracil in RNA and further uracil in DNA or thymine in RNA is considered a modified nucleotide like nucleotides.
Amino Acid
Now we will discuss about presentation of amino acids. The amino acids in an amino acid sequence must be represented in the amino two carboxy direction, like the five prime to three prime direction and the amino and carboxyl groups, they must not be shown in this sequence.
Regarding numbering, the first amino acid in this sequence is residue position number one, including amino acid preceding the mature protein like pre sequences and signal sequences.
And in case of circular amino acid sequences, applicant must choose the amino acid in residue position number one and number is continuous through the entire sequence, like we discussed in the nucleotide sequence presentation. Also the most restrictive symbol should be used.
For example, we have espartic acid or asparagne the symbol B should be used rather than X. The symbol X will be construed as any one of the amino acids coming to presentation of modified amino acids.Amino acid sequences separated by internal terminatorsignals, symbols like ter or s tree. They must be included as separate sequences because they are terminators. So that means the sequence is ended and we need to represent those sequences separately, giving them individual sequence identification numbers.
Modified Amino Acid
Further, the modified amino acids including the amino acids should be represented in this sequence as the corresponding unmodified amino acids whenever possible. And any modified amino acid in a sequence that cannot be otherwise represented by any other symbol, they should be presented with the symbol X.
Like we represent nucleotides by the symbol N and they should be further described in the feature table with feature keys and feature qualifiers.
The format for presenting amino acid sequences include all single letter upper case symbols, no spaces and no numbering should be included. And X has a default value of any one of the amino acids.
Tools for preparing Sequence Listing Standard
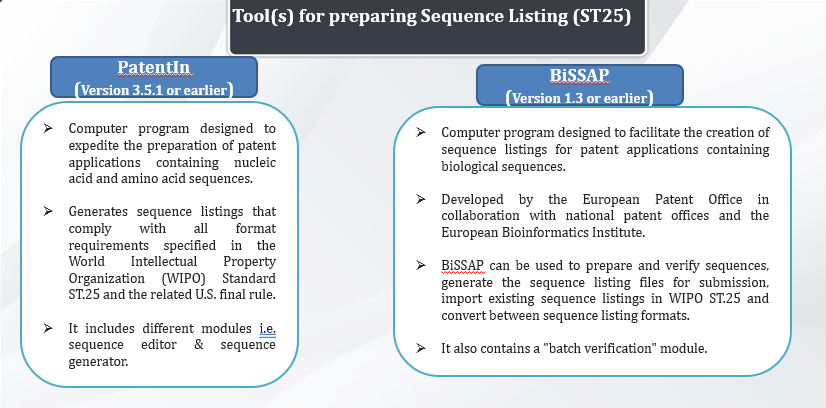

Preparing as per St 25 Sequence Listing Standard
Most commonly used softwares are patentIN and BiSSAP, wherein patentin is developed by USPTO and BiSSAP is developed by European Patent Office in collaboration with National Patent Offices and the European Bioinformatics Institute. Since they are currently used for St 25 sequences, we won’t discuss them in much detail.
Preparing as per St 26 Sequence Listing Standard
So to prepare St 26 compliant sequences, WIPO has developed WIPO sequence software to support authoring validation and generation of sequence listings.
WIPO Software For Sequence Listing
Member states requested that WIPO developed this common tool for all offices and applicants at international, national and regional level.
Further, the use of WIPO sequence software simplifies Stories XML creation with the user friendly interface wherein there is no need to ever directly edit an XML file, the sequence information can be saved in a project validated and then a sequence listing in St 25 26 format can be generated that is in the form of XML file.
Further, the data can be imported from this is very important that we can import data on WIPO sequence tool from St 26 sequence listings projects in St 25 sequence listings, multi sequence format files and the other kind of files. Software allows the validation of sequence listings in XML format as well. So this is the interface of WIPO sequence software. So when we open the WIPO sequence software, this is the front page that you’ll see, wherein we can see the tab new Project wherein we can start preparing a new sequence listing by naming it first and then entering our sequences.
Understanding to use the software
These are the two options import project and import sequence listings wherein we can import other files and then we can work on them further. The next tab is to validate sequence listing. Once we prepare our sequence listings, we can validate by using this tab. This information is present under the project header and here this project name applicant files. Your all previous projects will be shown here and you can click on them and work on them further if you wish. Then we can also enter persons and Organization information or Organism information from here. So this is the front page.
General Information Section
Next is the general information section when we click on the sequence, when we start entering our sequences. This is the first page, the general information wherein we enter the bibliographic information related to the patent application. We can see the headers here application identified, application filed, the applicant file reference number, filing date, application number, the priority application information can be added here. Then there’s IP Office application number, filing date, applicant inventors can be added here. Then we have a tab for title of the invention.
Sequence Section
So this was about general information section. Next we have sequences section wherein we enter our sequence and its details. The sequence is entered here, then w have a sequence ID numbers, its name then length is it automatically picked up. Molecular type is DNA, then organism. We need to select it and then once we select the information, it is presented in this manner. Then in case of modified sequences, there are feature keys to add the feature and qualifiers. Earlier when we used to pay tint in, we used to prepare sequence listing in St 25 format. This was the format we used to receive the sequence listings in however, XML format sequences looks like this. This is the XML file that will be generated once all the information of sequences entered.
I know it is not easy to understand by its personal look, but bipose sequence also gives an option of HTML format that is relatively easy to read.
Sequence Listing Standard- Elements of the Sequence
Now let’s discuss the sequence elements of this XML file one by one. I have listed down the sequence elements here also because it is difficult to read and understand from here. But still we will try to read here.
The first sequence element is application identification and this header is composed of IP Office code IB . And then application number text like here is the application number and the IP Office code is IB.
Then filing date is also given where we can enter the filing date. Then we have applicant reference number. This applicant file reference, it is a single unique identifier assigned by applicant to identify a particular application. Then we have earliest priority application identification here wherein the details are again the IP Office application number and filing date.
After that we have the applicant or assignee name which is shown here, the invention title and after that then we also have length, small type, the feature locations and organism names, the feature keys and feature qualifiers.
So let’s see the sequence element details here also these are the bibliographic information and these are the sequence information. And for modified sequences the feature key, feature location, feature qualifiers are used.
Feature keys and Qualifiers
So to use the feature keys and qualifiers in the best possible way. This is one example that I’ve taken that in case of naturally occurring mutations. We can select the feature key as variation for nucleic assets and inthe qualifier we can use replace or note. And similarly we have other examples. This is the table that is differentiates the feature keys. Like in St 25 we used to have the feature key as allele but in St 26 the feature key is used as miscellaneous feature. Then under qualifier we specify it as allele and similarly there are a lot of examples that are given in St 26 document now that clearly states the differences between the two.
Understanding with the help of an Example
Let’s take an example here the live example so that we can understand what we discussed earlier.
So this is a sequence, the ten nucleotide sequence Gagcatttac- AP-taaggct. Then this wherein AP is an a basic site. So should the sequence be listed? Well, yes, the sequence should be listed because the number of nucleotides is ten or more than ten in this sequence.
So we need to list this and we will consider the AP as N. So the sequence would be presented as the first stretch of nucleotide sequence.
Then N in place of then the second stretch of nucleotides and in the annotations. We will explain that feature key is modified base. Feature qualifier as mod base then under the qualifier value will enter other. We’ll write a note that would include and describe the modified base as an Abasic site.
Another Example
Let’s take another example of a branch to amino acid sequence. As you can see here, we have one
strand on the top and then the Lysine is connected with Glycine forming another branch.
So should the sequence be listed? Yes, it should be listed because the unbranched or linear region of a sequence containing four or more specifically defined amino acids must be included in a sequence listing.
ow, how should the sequence be presented? Now, it will be divided into two strands.
We’ll pick the first till Lysine and then we’ll pick the second strand. And both these sequences will be provided with separate sequence identifiers.
One would be given one and two, the annotations feature key would be same for both.
And then we’ll give a note in feature qualifier with these comments.
But now you might have a question. Why did we use this stretch? Lysine lysine lysine because the number of amino acids is less than four. So we will not include us. That’s all. We have included this already. Moving forward, we have reached the end of the Webinar. What we have discussed in the Webinar is shown here. You can just go through these and refresh what we discussed in this Webinar. I’m happy to take questions if you have any.
Questions and Answers
Devica is speaking – Thank you, Pooja. This has been quite an informative session. We do have some questions first thing regarding this PPT. So participants who are interested, they can drop us an email at Webinar@sagaciousresearch.com. If they have any further questions or if they would want more detailed support from our side. Apart from that, there is one question.
When will the standard come into effect?
Pooja is speaking – Well, the standard is set to come into effect on 1st July 22, if the date is not postponed further.
How can previously generated St 25 sequence listings be most easily converted to St 26 format?
Pooja is speaking – The WIPO sequence software, it provides a feature of importing St 25 files on its software. So we can convert St 25 using that. But let me mention that we still need to make few changes manually. After we do that, we can extract the file in XML format. This is the easiest way. We import the SD 25 file on WIPO sequence and then convert it into XML. Okay.
How does this apply at USPTO or other PTO?
Pooja is speaking – Well, as per the WIPO guidelines, all patent offices, including USPTO, should simultaneously transit at the Pct national and regional levels.
End Note
Devica is speaking – Okay, so there are a few questions that could not be taken up due to time limit. We have reached past the time limit. We hope to cover them through our subsequent write up, which we will publish, post this webinar. And once again, I would say that participants can drop us an email at webinar@sagaciousresearch.com. So this has been a wonderful session overall.
I’m sure our listeners have great takeaways from this session. They will be able to use several pointers when working on sequence listings. I want to extend a big thanks to our listeners who helped us start on task time and finish. Highly appreciate that and thank you very much. Please join us in our next webinar. Thank you, Pooja, for joining us. Have a great day ahead. Thank you, everyone. Bye.




